Porcine reproductive and respiratory syndrome (PRRS) and porcine epidemic diarrhea (PED) cost the US swine industry more than $1 billion/year. Our goals combine data-visualization with spatial and statistical methods to increase the fundamental knowledge about PED and PRRS spatial epidemiology, enhancing our understanding of disease spread to develop novel data-integration tools to optimize disease control.
Publications
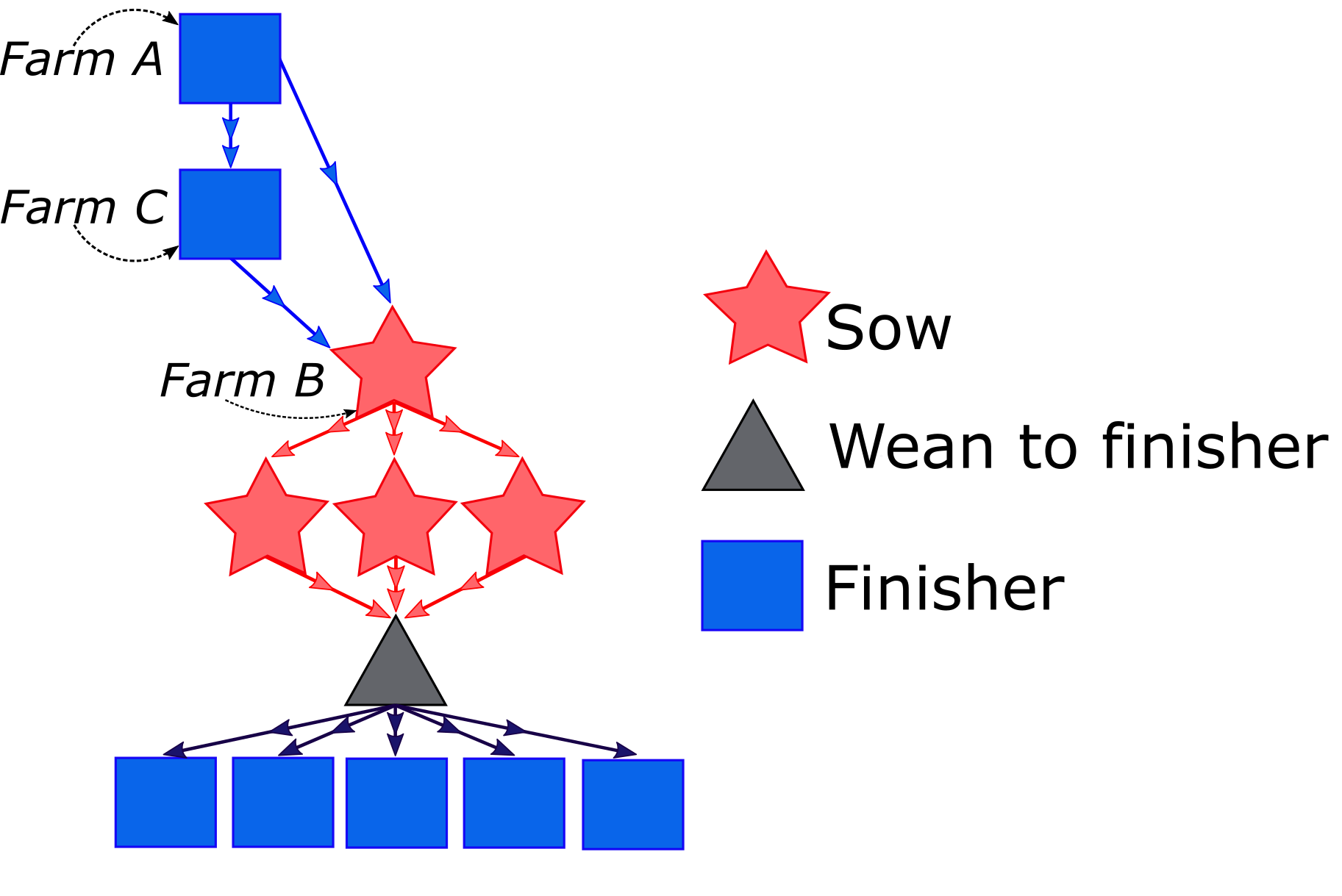
Descriptive and risk analysis of vehicle movements linked to porcine reproductive and respiratory syndrome and porcine epidemic diarrhea transmission in US commercial swine farms

Truck Cleaning and Disinfection, and the Risk of PRRSV Dissemination in Multi-Site Pig Production Systems in the United States
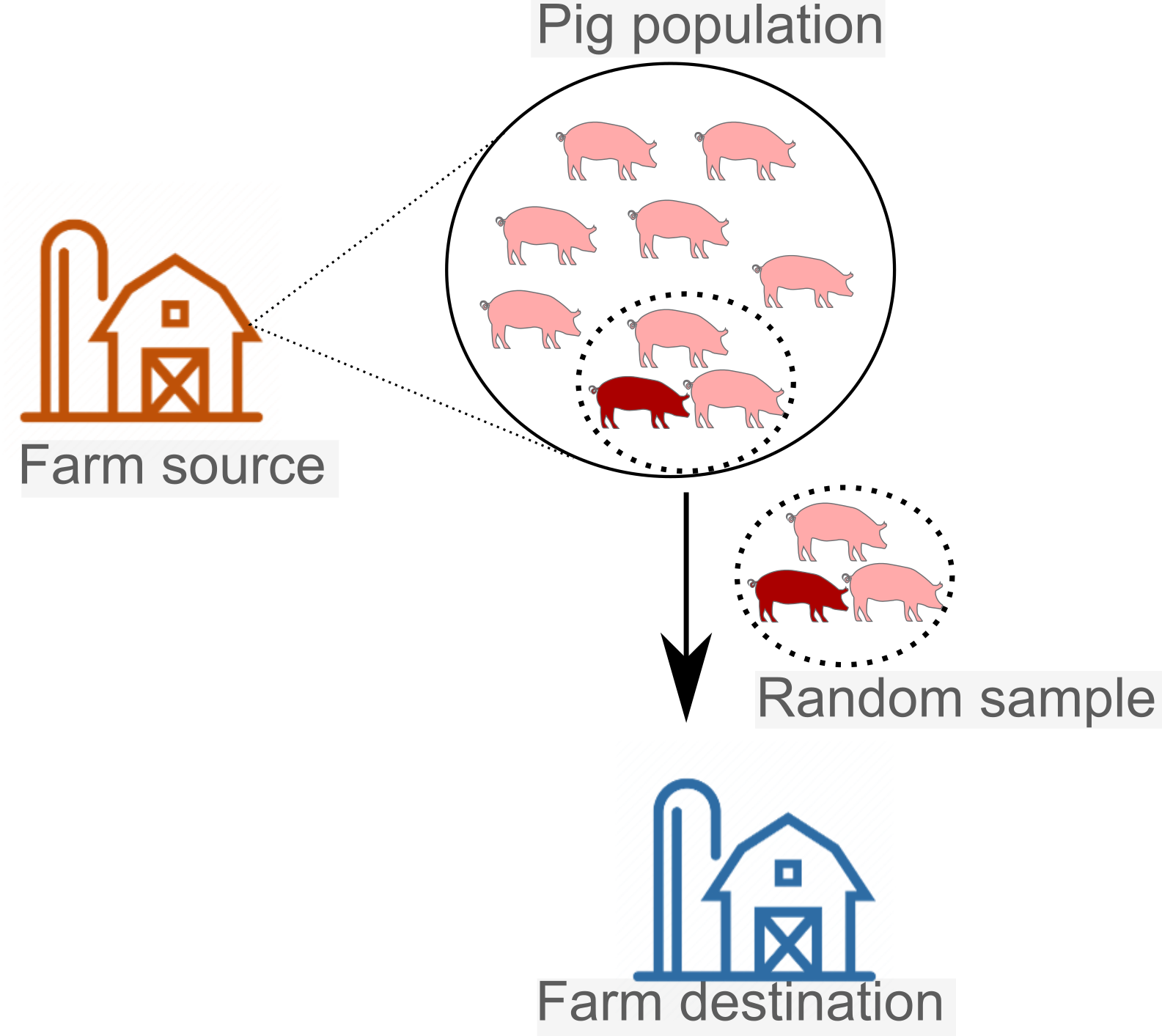
Evaluation of porcine epidemic diarrhea virus RNA contamination on swine industry transportation vehicles

Modeling between-farm transmission dynamics of porcine epidemic diarrhea virus: characterizing the dominant transmission routes

A discrete-time survival model for porcine epidemic diarrhea virus

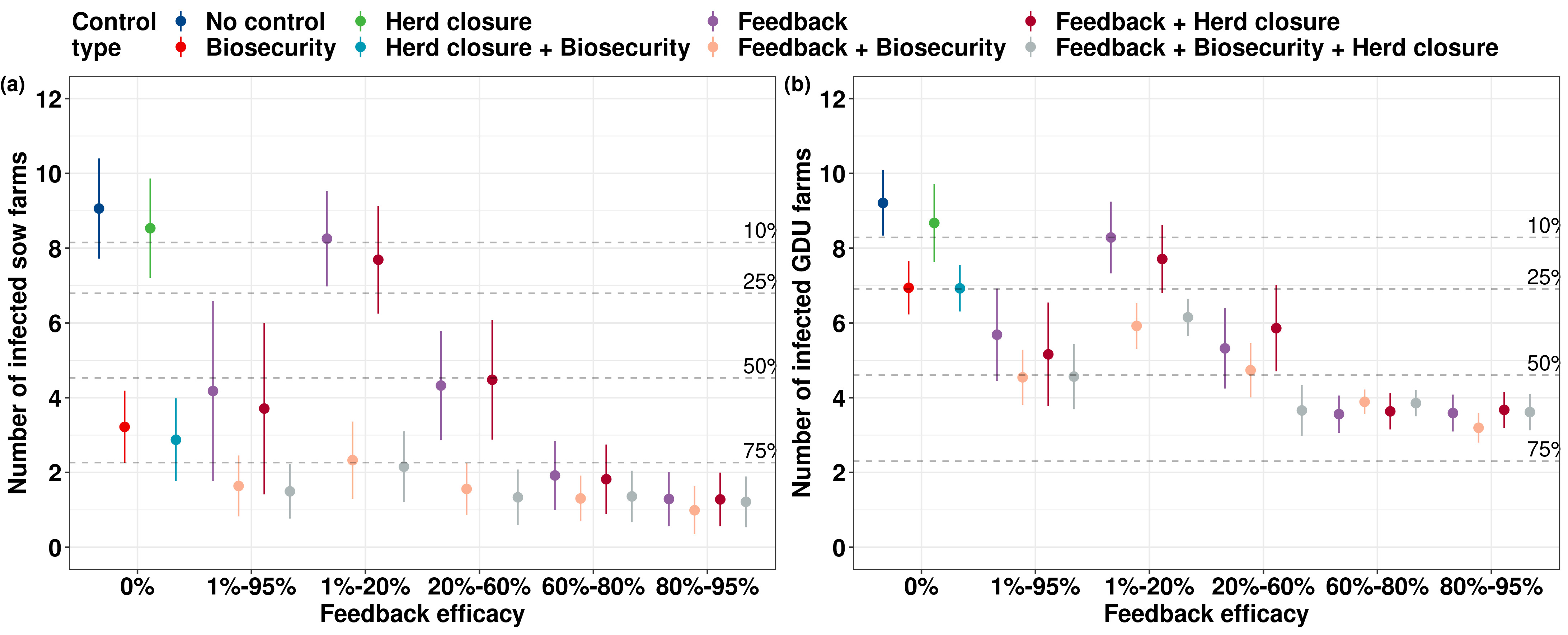
The between-farm transmission dynamics of Porcine Epidemic Diarrhea Virus: A short-term forecast modeling comparison and the effectiveness of control strategies
Porcine reproductive and respiratory syndrome virus dissemination across pig production systems in the United States

Identifying outbreaks of Porcine Epidemic Diarrhea virus through animal movements and spatial neighborhoods
Funding
We are grateful for the funding agencies:
